Partial dependence / Individual Conditional Expectation (PD/ICE) Explainer Demo
This example demonstrates how to interpret a scikit-learn model using the H2O Sonar library and retrieve the data and partial dependence plot.
[1]:
import logging
import os
import pandas
import datatable
import webbrowser
from h2o_sonar import interpret
from h2o_sonar.lib.api import commons, explainers
from h2o_sonar.lib.api.models import ModelApi
from h2o_sonar.explainers.pd_ice_explainer import PdIceExplainer
from sklearn.ensemble import GradientBoostingClassifier
[2]:
# dataset
dataset_path = "../../data/predictive/creditcard.csv"
target_col = "default payment next month"
df = pandas.read_csv(dataset_path)
(X, y) = df.drop(target_col, axis=1), df[target_col]
# results
results_location = "../../results"
os.makedirs(results_location, exist_ok=True)
[3]:
# explainer description
interpret.describe_explainer(PdIceExplainer)
[3]:
{'id': 'h2o_sonar.explainers.pd_ice_explainer.PdIceExplainer',
'name': 'PdIceExplainer',
'display_name': 'Partial Dependence Plot',
'tagline': 'PdIceExplainer.',
'description': 'Partial dependence plot (PDP) portrays the average prediction\nbehavior of the model across the domain of an input variable along with +/- 1\nstandard deviation bands. Individual Conditional Expectations plot (ICE) displays\nthe prediction behavior for an individual row of data when an input variable is\ntoggled across its domain.\n\nPD binning:\n\n**Integer** feature:\n\n* bins in **numeric** mode:\n * bins are integers\n * (at most) `grid_resolution` integer values in between minimum and maximum\n of feature values\n * bin values are created as evenly as possible\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are integers\n * top `grid_resolution` values from feature values ordered by frequency\n (int values are converted to strings and most frequent values are used\n as bins)\n* quantile bins in **numeric** mode:\n * bins are integers\n * bin values are created with the aim that there will be the same number of\n observations per bin\n * q-quantile used to created ``q`` bins where ``q`` is specified by PD parameter\n* quantile bins in **categorical** mode:\n * not supported\n\n**Float** feature:\n\n* bins in **numeric** mode:\n * bins are floats\n * `grid_resolution` float values in between minimum and maximum of feature\n values\n * bin values are created as evenly as possible\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are floats\n * top `grid_resolution` values from feature values ordered by frequency\n (float values are converted to strings and most frequent values are used\n as bins)\n* quantile bins in **numeric** mode:\n * bins are floats\n * bin values are created with the aim that there will be the same number of\n observations per bin\n * q-quantile used to created ``q`` bins where ``q`` is specified by PD parameter\n* quantile bins in **categorical** mode:\n * not supported\n\n**String** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * bins are strings\n * top `grid_resolution` values from feature values ordered by frequency\n* quantile bins:\n * not supported\n\n**Date/datetime** feature:\n\n* bins in **numeric** mode:\n * bins are dates\n * `grid_resolution` date values in between minimum and maximum of feature\n values\n * bin values are created as evenly as possible:\n 1. dates are parsed and converted to epoch timestamps i.e integers\n 2. bins are created as in case of numeric integer binning\n 3. integer bins are converted back to original date format\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are dates\n * top `grid_resolution` values from feature values ordered by frequency\n (dates are handled as opaque strings and most frequent values are used\n as bins)\n* quantile bins:\n * not supported\n\nPD out of range binning:\n\n**Integer** feature:\n\n* OOR bins in **numeric** mode:\n * OOR bins are integers\n * (at most) `oor_grid_resolution` integer values are added below minimum and\n above maximum\n * bin values are created by adding/substracting rounded standard deviation\n (of feature values) above and below maximum and minimum `oor_grid_resolution`\n times\n * 1 used used if rounded standard deviation would be 0\n * if feature is of unsigned integer type, then bins below 0\n are not created\n * if rounded standard deviation and/or `oor_grid_resolution` is so high\n that it would cause lower OOR bins to be negative numbers, then standard\n deviation of size 1 is tried instead\n* OOR bins in **categorical** mode:\n * same as numeric mode\n\n**Float** feature:\n\n* OOR bins in **numeric** mode:\n * OOR bins are floats\n * `oor_grid_resolution` float values are added below minimum and above maximum\n * bin values are created by adding/substracting standard deviation\n (of feature values) above and below maximum and minimum `oor_grid_resolution`\n times\n* OOR bins in **categorical** mode:\n * same as numeric mode\n\n**String** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * OOR bins are strings\n * value `UNSEEN` is added as OOR bin\n\n**Date** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * OOR bins are strings\n * value `UNSEEN` is added as OOR bin\n\n',
'brief_description': 'PdIceExplainer.',
'model_types': ['iid', 'time_series'],
'can_explain': ['regression', 'binomial', 'multinomial'],
'explanation_scopes': ['global_scope', 'local_scope'],
'explanations': [{'explanation_type': 'global-partial-dependence',
'name': 'PartialDependenceExplanation',
'category': '',
'scope': 'global',
'has_local': '',
'formats': []},
{'explanation_type': 'local-individual-conditional-explanation',
'name': 'IndividualConditionalExplanation',
'category': '',
'scope': 'local',
'has_local': '',
'formats': []}],
'keywords': ['run-by-default',
'can-add-feature',
'explains-feature-behavior',
'h2o-sonar'],
'parameters': [{'name': 'sample_size',
'description': 'Sample size for Partial Dependence Plot.',
'comment': '',
'type': 'int',
'val': 25000,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'max_features',
'description': 'Partial Dependence Plot number of features (to see all features used by model set to -1).',
'comment': '',
'type': 'int',
'val': 10,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'features',
'description': 'Partial Dependence Plot feature list.',
'comment': '',
'type': 'list',
'val': None,
'predefined': [],
'tags': ['SOURCE_DATASET_COLUMN_NAMES'],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'oor_grid_resolution',
'description': 'Partial Dependence Plot number of out of range bins.',
'comment': '',
'type': 'int',
'val': 0,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'quantile-bin-grid-resolution',
'description': 'Partial Dependence Plot quantile binning (total quantile points used to create bins).',
'comment': '',
'type': 'int',
'val': 0,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'grid_resolution',
'description': 'Partial Dependence Plot observations per bin (number of equally spaced points used to create bins).',
'comment': '',
'type': 'int',
'val': 20,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'center',
'description': 'Center Partial Dependence Plot using ICE centered at 0.',
'comment': '',
'type': 'bool',
'val': False,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'sort_bins',
'description': 'Ensure bin values sorting.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'histograms',
'description': 'Enable histograms.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'quantile-bins',
'description': 'Per-feature quantile binning (Example: if choosing features\n F1 and F2, this parameter is \'{"F1": 2,"F2": 5}\'. Note, you can\n set all features to use the same quantile binning with the\n `Partial Dependence Plot quantile binning` parameter and then\n adjust the quantile binning for a subset of PDP features with\n this parameter).',
'comment': '',
'type': 'str',
'val': '',
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'numcat_num_chart',
'description': 'Unique feature values count driven Partial Dependence Plot binning and chart selection.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'numcat_threshold',
'description': 'Threshold for Partial Dependence Plot binning and chart selection (<=threshold categorical, >threshold numeric).',
'comment': '',
'type': 'int',
'val': 11,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'debug_residuals',
'description': 'Debug model residuals.',
'comment': '',
'type': 'bool',
'val': False,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''}],
'metrics_meta': []}
Interpretation
[4]:
# scikit-learn model
gradient_booster = GradientBoostingClassifier(learning_rate=0.1)
gradient_booster.fit(X, y)
# explainable model
model = ModelApi().create_model(target_col=target_col, model_src=gradient_booster, used_features=X.columns.to_list())
interpretation = interpret.run_interpretation(
dataset=df,
model=model,
target_col=target_col,
results_location=results_location,
log_level=logging.INFO,
explainers=[
commons.ExplainerToRun(
explainer_id=PdIceExplainer.explainer_id(),
params="",
)
]
)
/home/user/h/mli/git/h2o-sonar-FLOSS/.venv/lib/python3.11/site-packages/ragas/metrics/__init__.py:1: LangChainDeprecationWarning: As of langchain-core 0.3.0, LangChain uses pydantic v2 internally. The langchain_core.pydantic_v1 module was a compatibility shim for pydantic v1, and should no longer be used. Please update the code to import from Pydantic directly.
For example, replace imports like: `from langchain_core.pydantic_v1 import BaseModel`
with: `from pydantic import BaseModel`
or the v1 compatibility namespace if you are working in a code base that has not been fully upgraded to pydantic 2 yet. from pydantic.v1 import BaseModel
from ragas.metrics._answer_correctness import AnswerCorrectness, answer_correctness
/home/user/h/mli/git/h2o-sonar-FLOSS/.venv/lib/python3.11/site-packages/ragas/metrics/__init__.py:4: LangChainDeprecationWarning: As of langchain-core 0.3.0, LangChain uses pydantic v2 internally. The langchain.pydantic_v1 module was a compatibility shim for pydantic v1, and should no longer be used. Please update the code to import from Pydantic directly.
For example, replace imports like: `from langchain.pydantic_v1 import BaseModel`
with: `from pydantic import BaseModel`
or the v1 compatibility namespace if you are working in a code base that has not been fully upgraded to pydantic 2 yet. from pydantic.v1 import BaseModel
from ragas.metrics._context_entities_recall import (
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
X does not have valid feature names, but GradientBoostingClassifier was fitted with feature names
2026-01-29 16:11:39,916 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 saving PD to ../../results/h2o-sonar/mli_experiment_ea8d1116-5f12-4d93-b0c6-91453d509ffd/explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-pd-dai-model.json
2026-01-29 16:11:39,918 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 computation finished & stored to: ../../results/h2o-sonar/mli_experiment_ea8d1116-5f12-4d93-b0c6-91453d509ffd/explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-pd-dai-model.json
2026-01-29 16:11:39,925 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: ID/True
2026-01-29 16:11:39,928 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: ID/False
2026-01-29 16:11:39,931 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: LIMIT_BAL/True
2026-01-29 16:11:39,932 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: LIMIT_BAL/False
2026-01-29 16:11:39,933 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: SEX/True
2026-01-29 16:11:39,934 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: SEX/False
2026-01-29 16:11:39,935 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: EDUCATION/True
2026-01-29 16:11:39,936 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: EDUCATION/False
2026-01-29 16:11:39,937 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: MARRIAGE/True
2026-01-29 16:11:39,938 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: MARRIAGE/False
2026-01-29 16:11:39,939 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: AGE/True
2026-01-29 16:11:39,940 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: AGE/False
2026-01-29 16:11:39,941 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_0/True
2026-01-29 16:11:39,943 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_0/False
2026-01-29 16:11:39,944 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_2/True
2026-01-29 16:11:39,947 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_2/False
2026-01-29 16:11:39,949 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_3/True
2026-01-29 16:11:39,950 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_3/False
2026-01-29 16:11:39,952 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_4/True
2026-01-29 16:11:39,953 - h2o_sonar.explainers.pd_ice_explainer.PdIceExplainerLogger - INFO - PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 creating histogram: PAY_4/False
Explainer Result
[5]:
# retrieve the result
result = interpretation.get_explainer_result(PdIceExplainer.explainer_id())
[6]:
# open interpretation HTML report in web browser
webbrowser.open(interpretation.result.get_html_report_location())
[6]:
True
[7]:
# summary
result.summary()
[7]:
{'id': 'h2o_sonar.explainers.pd_ice_explainer.PdIceExplainer',
'name': 'PdIceExplainer',
'display_name': 'Partial Dependence Plot',
'tagline': 'PdIceExplainer.',
'description': 'Partial dependence plot (PDP) portrays the average prediction\nbehavior of the model across the domain of an input variable along with +/- 1\nstandard deviation bands. Individual Conditional Expectations plot (ICE) displays\nthe prediction behavior for an individual row of data when an input variable is\ntoggled across its domain.\n\nPD binning:\n\n**Integer** feature:\n\n* bins in **numeric** mode:\n * bins are integers\n * (at most) `grid_resolution` integer values in between minimum and maximum\n of feature values\n * bin values are created as evenly as possible\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are integers\n * top `grid_resolution` values from feature values ordered by frequency\n (int values are converted to strings and most frequent values are used\n as bins)\n* quantile bins in **numeric** mode:\n * bins are integers\n * bin values are created with the aim that there will be the same number of\n observations per bin\n * q-quantile used to created ``q`` bins where ``q`` is specified by PD parameter\n* quantile bins in **categorical** mode:\n * not supported\n\n**Float** feature:\n\n* bins in **numeric** mode:\n * bins are floats\n * `grid_resolution` float values in between minimum and maximum of feature\n values\n * bin values are created as evenly as possible\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are floats\n * top `grid_resolution` values from feature values ordered by frequency\n (float values are converted to strings and most frequent values are used\n as bins)\n* quantile bins in **numeric** mode:\n * bins are floats\n * bin values are created with the aim that there will be the same number of\n observations per bin\n * q-quantile used to created ``q`` bins where ``q`` is specified by PD parameter\n* quantile bins in **categorical** mode:\n * not supported\n\n**String** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * bins are strings\n * top `grid_resolution` values from feature values ordered by frequency\n* quantile bins:\n * not supported\n\n**Date/datetime** feature:\n\n* bins in **numeric** mode:\n * bins are dates\n * `grid_resolution` date values in between minimum and maximum of feature\n values\n * bin values are created as evenly as possible:\n 1. dates are parsed and converted to epoch timestamps i.e integers\n 2. bins are created as in case of numeric integer binning\n 3. integer bins are converted back to original date format\n * minimum and maximum is included in bins\n (if `grid_resolution` is bigger or equal to 2)\n* bins in **categorical** mode:\n * bins are dates\n * top `grid_resolution` values from feature values ordered by frequency\n (dates are handled as opaque strings and most frequent values are used\n as bins)\n* quantile bins:\n * not supported\n\nPD out of range binning:\n\n**Integer** feature:\n\n* OOR bins in **numeric** mode:\n * OOR bins are integers\n * (at most) `oor_grid_resolution` integer values are added below minimum and\n above maximum\n * bin values are created by adding/substracting rounded standard deviation\n (of feature values) above and below maximum and minimum `oor_grid_resolution`\n times\n * 1 used used if rounded standard deviation would be 0\n * if feature is of unsigned integer type, then bins below 0\n are not created\n * if rounded standard deviation and/or `oor_grid_resolution` is so high\n that it would cause lower OOR bins to be negative numbers, then standard\n deviation of size 1 is tried instead\n* OOR bins in **categorical** mode:\n * same as numeric mode\n\n**Float** feature:\n\n* OOR bins in **numeric** mode:\n * OOR bins are floats\n * `oor_grid_resolution` float values are added below minimum and above maximum\n * bin values are created by adding/substracting standard deviation\n (of feature values) above and below maximum and minimum `oor_grid_resolution`\n times\n* OOR bins in **categorical** mode:\n * same as numeric mode\n\n**String** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * OOR bins are strings\n * value `UNSEEN` is added as OOR bin\n\n**Date** feature:\n\n* bins in **numeric** mode:\n * not supported\n* bins in **categorical** mode:\n * OOR bins are strings\n * value `UNSEEN` is added as OOR bin\n\n',
'brief_description': 'PdIceExplainer.',
'model_types': ['iid', 'time_series'],
'can_explain': ['regression', 'binomial', 'multinomial'],
'explanation_scopes': ['global_scope', 'local_scope'],
'explanations': [{'explanation_type': 'global-partial-dependence',
'name': 'Partial Dependence Plot (PDP)',
'category': 'DAI MODEL',
'scope': 'global',
'has_local': 'local-individual-conditional-explanation',
'formats': ['application/json']},
{'explanation_type': 'local-individual-conditional-explanation',
'name': 'Individual Conditional Expectations (ICE)',
'category': 'DAI MODEL',
'scope': 'local',
'has_local': None,
'formats': ['application/vnd.h2oai.json+datatable.jay']},
{'explanation_type': 'global-html-fragment',
'name': 'Partial Dependence Plot (PDP)',
'category': 'DAI MODEL',
'scope': 'global',
'has_local': None,
'formats': ['text/html']}],
'keywords': ['run-by-default',
'can-add-feature',
'explains-feature-behavior',
'h2o-sonar'],
'parameters': [{'name': 'sample_size',
'description': 'Sample size for Partial Dependence Plot.',
'comment': '',
'type': 'int',
'val': 25000,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'max_features',
'description': 'Partial Dependence Plot number of features (to see all features used by model set to -1).',
'comment': '',
'type': 'int',
'val': 10,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'features',
'description': 'Partial Dependence Plot feature list.',
'comment': '',
'type': 'list',
'val': None,
'predefined': [],
'tags': ['SOURCE_DATASET_COLUMN_NAMES'],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'oor_grid_resolution',
'description': 'Partial Dependence Plot number of out of range bins.',
'comment': '',
'type': 'int',
'val': 0,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'quantile-bin-grid-resolution',
'description': 'Partial Dependence Plot quantile binning (total quantile points used to create bins).',
'comment': '',
'type': 'int',
'val': 0,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'grid_resolution',
'description': 'Partial Dependence Plot observations per bin (number of equally spaced points used to create bins).',
'comment': '',
'type': 'int',
'val': 20,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'center',
'description': 'Center Partial Dependence Plot using ICE centered at 0.',
'comment': '',
'type': 'bool',
'val': False,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'sort_bins',
'description': 'Ensure bin values sorting.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'histograms',
'description': 'Enable histograms.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'quantile-bins',
'description': 'Per-feature quantile binning (Example: if choosing features\n F1 and F2, this parameter is \'{"F1": 2,"F2": 5}\'. Note, you can\n set all features to use the same quantile binning with the\n `Partial Dependence Plot quantile binning` parameter and then\n adjust the quantile binning for a subset of PDP features with\n this parameter).',
'comment': '',
'type': 'str',
'val': '',
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'numcat_num_chart',
'description': 'Unique feature values count driven Partial Dependence Plot binning and chart selection.',
'comment': '',
'type': 'bool',
'val': True,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'numcat_threshold',
'description': 'Threshold for Partial Dependence Plot binning and chart selection (<=threshold categorical, >threshold numeric).',
'comment': '',
'type': 'int',
'val': 11,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''},
{'name': 'debug_residuals',
'description': 'Debug model residuals.',
'comment': '',
'type': 'bool',
'val': False,
'predefined': [],
'tags': [],
'min_': 0.0,
'max_': 0.0,
'category': ''}],
'metrics_meta': []}
[8]:
# parameters
result.params()
[8]:
{'sample_size': 25000,
'max_features': 10,
'features': None,
'oor_grid_resolution': 0,
'quantile-bin-grid-resolution': 0,
'grid_resolution': 20,
'center': False,
'sort_bins': True,
'histograms': True,
'quantile-bins': '',
'numcat_num_chart': True,
'numcat_threshold': 11,
'debug_residuals': False}
Display PD Data
[9]:
result.data(feature_name="EDUCATION")
[9]:
| bin | frequency | pd | sd | oor | |
|---|---|---|---|---|---|
| ▪▪▪▪ | ▪▪▪▪ | ▪▪▪▪▪▪▪▪ | ▪▪▪▪▪▪▪▪ | ▪ | |
| 0 | 0 | 3 | 0.2227 | 0.416079 | 0 |
| 1 | 1 | 3714 | 0.2227 | 0.416079 | 0 |
| 2 | 2 | 4588 | 0.3414 | 0.474203 | 0 |
| 3 | 3 | 1590 | 0.3414 | 0.474203 | 0 |
| 4 | 4 | 26 | 0.3414 | 0.474203 | 0 |
| 5 | 5 | 79 | 0.2298 | 0.420725 | 0 |
| 6 | 6 | NA | 0.2298 | 0.420725 | 0 |
[10]:
result.data(feature_name="PAY_3")
[10]:
| bin | frequency | pd | sd | oor | |
|---|---|---|---|---|---|
| ▪▪▪▪ | ▪▪▪▪ | ▪▪▪▪▪▪▪▪ | ▪▪▪▪▪▪▪▪ | ▪ | |
| 0 | −2 | 1292 | 0.3048 | 0.460346 | 0 |
| 1 | −1 | 2143 | 0.3048 | 0.460346 | 0 |
| 2 | 0 | 5125 | 0.3048 | 0.460346 | 0 |
| 3 | 1 | 3 | 0.3271 | 0.469178 | 0 |
| 4 | 2 | 1306 | 0.326 | 0.468771 | 0 |
| 5 | 3 | 58 | 0.3259 | 0.468734 | 0 |
| 6 | 4 | 31 | 0.3259 | 0.468734 | 0 |
| 7 | 5 | 13 | 0.3259 | 0.468734 | 0 |
| 8 | 6 | 12 | 0.3259 | 0.468734 | 0 |
| 9 | 7 | 17 | 0.3259 | 0.468734 | 0 |
| 10 | 8 | NA | 0.3259 | 0.468734 | 0 |
Plot PD Data
[11]:
result.plot(feature_name="EDUCATION", override_feature_type=result.format.KEY_CATEGORICAL)
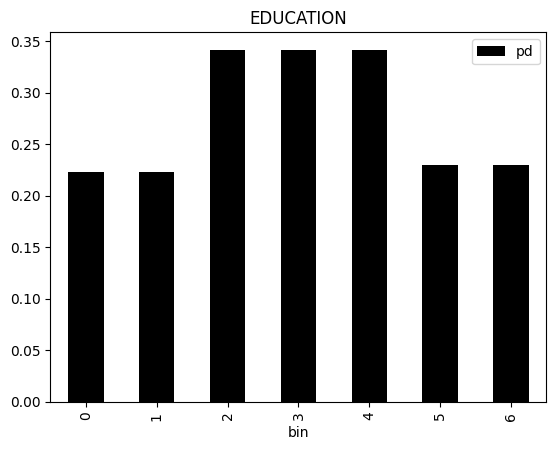
[12]:
result.plot(feature_name="PAY_3")
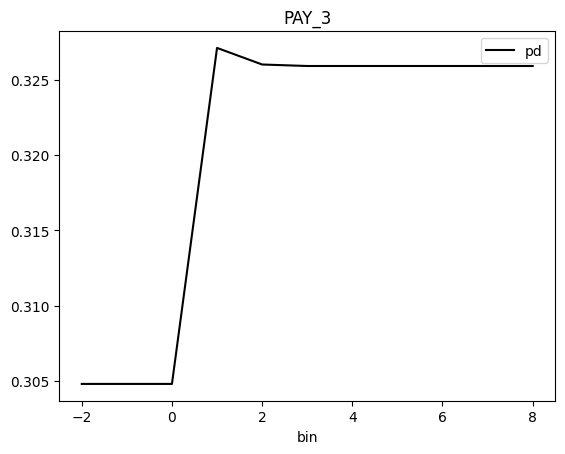
Save Explainer Log and Data
[13]:
# save the explainer log
result.log(path="./pd-ice-demo.log")
[14]:
!head pd-ice-demo.log
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 BEGIN calculation
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 loading dataset
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 loaded dataset has 10000 rows and 25 columns
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 getting features list, importance and metadata
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 all most important model features: ['ID', 'LIMIT_BAL', 'SEX', 'EDUCATION', 'MARRIAGE', 'AGE', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6', 'BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6', 'PAY_AMT1', 'PAY_AMT2', 'PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6']
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 features used by model: ['ID', 'LIMIT_BAL', 'SEX', 'EDUCATION', 'MARRIAGE', 'AGE', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6', 'BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6', 'PAY_AMT1', 'PAY_AMT2', 'PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6']
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25: calculating PD for features ['ID', 'LIMIT_BAL', 'SEX', 'EDUCATION', 'MARRIAGE', 'AGE', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4']
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 feature metadata: {'id': [], 'categorical': [], 'numeric': [], 'catnum': [], 'date': [], 'time': [], 'datetime': [], 'text': [], 'image': [], 'date-format': [], 'quantile-bin': {}}
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 1 frame strategy: True
2026-01-29 16:11:38,082 INFO PD/ICE ea8d1116-5f12-4d93-b0c6-91453d509ffd/cba719fb-5061-483d-add5-79c4175d2b25 residual PD/ICE should NOT be calculated, but y has been specified - setting it None
[15]:
# save the explainer data
result.zip(file_path="./pd-ice-demo-archive.zip")
[16]:
!unzip -l pd-ice-demo-archive.zip
Archive: pd-ice-demo-archive.zip
Length Date Time Name
--------- ---------- ----- ----
11536 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/result_descriptor.json
151 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json.meta
6978 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/explanation.json
929 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_3_class_0.json
2557 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_5_class_0.json
321 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_2_class_0.json
2664 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_0_class_0.json
1422 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_8_class_0.json
1411 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_6_class_0.json
1410 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_9_class_0.json
1413 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_7_class_0.json
2764 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_1_class_0.json
563 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_partial_dependence/application_json/pd_feature_4_class_0.json
2 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/problems/problems_and_actions.json
165 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay.meta
7080 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/explanation.json
1601936 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_0_class_0.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_8_class_0.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_9_class_0.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_7_class_0.jay
1601944 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_1_class_0.jay
560688 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_3_class_0.jay
320440 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_4_class_0.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_6_class_0.jay
1601784 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_5_class_0.jay
80184 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/y_hat.jay
160272 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/local_individual_conditional_explanation/application_vnd_h2oai_json_datatable_jay/ice_feature_2_class_0.jay
110 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html.meta
13559 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-6-class-0.png
13628 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-9-class-0.png
13571 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-2-class-0.png
15041 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-8-class-0.png
3707 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/explanation.html
14612 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-4-class-0.png
17683 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-1-class-0.png
19503 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-3-class-0.png
13504 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-7-class-0.png
18093 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-5-class-0.png
18934 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/global_html_fragment/text_html/pd-feature-0-class-0.png
5747 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/log/explainer_run_cba719fb-5061-483d-add5-79c4175d2b25.log
320440 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-5.jay
13618 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-pd-dai-model.json
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-8.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-9.jay
1601936 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-1.jay
560688 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-4.jay
1601784 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-6.jay
1601944 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-2.jay
80184 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/mli_dataset_y_hat.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-10.jay
881024 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-7.jay
160272 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model-3.jay
2349 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/work/h2o_sonar-ice-dai-model.json
2 2026-01-29 16:11 explainer_h2o_sonar_explainers_pd_ice_explainer_PdIceExplainer_cba719fb-5061-483d-add5-79c4175d2b25/insights/insights_and_actions.json
--------- -------
19127715 54 files
[ ]: